Upcoming training events for bioinformatics and genomics researchers
Between September and the end of 2022, NeSI and Genomics Aotearoa are co-hosting a range of hands-on skill-building events for Aotearoa New Zealand's bioinformatics and genomics communities. Follow the links in the 'Upcoming events' list below for more information or to register.
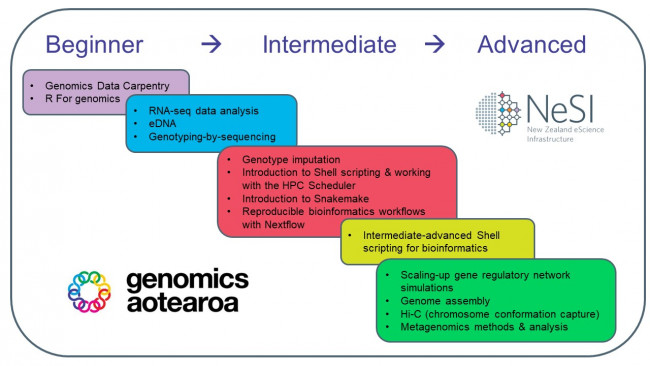
Also, the NeSI and Genomics Aotearoa training teams have developed a new course overview of their training programme in bioinformatics (a snapshot of which is pictured in the image below). Browse the offerings here. This programme is free for all participants. If you have any questions about the overall programme, upcoming sessions, or suggestions / requests for training in other areas of bioinformatics or genomics, contact training@nesi.org.nz.
Upcoming events
Intermediate-Advanced Shell for bioinformatics
Online
15 September 2022
Registration has closed
This online workshop includes:
- An overview of the Shell, UNIX and Linux
- Downloading data from a remote source and checking data integrity
- Recap navigating files and directories, and commands used in routine tasks
- Inspecting and manipulating data, part 1 (the head, less, grep, and sed commands)
- Inspecting and manipulating data, part 2 (using awk and bioawk to process text)
- Automating file processing
- Challenges: solve example molecular biology problems using shell scripts.
Scaling gene regulatory networks
Online
21-22 September 2022
9 am – 4 pm (each day)
Register here:
https://www.eventbrite.co.nz/e/scaling-gene-regulatory-networks-simulations-tickets-407150697697
This two-day workshop has been jointly developed by Olivia Angelin-Bonnet from Plant & Food Research, and the training team at NeSI and Genomics Aotearoa. Olivia researches on how to decipher genotype-phenotype interactions through regulatory networks using omics datasets, and will speak about simulating gene regulatory networks, using R and Julia. Some elaboration on what Olivia and Dini will focus in the workshop:
- Why simulations are valuable in systems biology,
- What regulatory networks are and how they can be modelled,
- How to use the sismonr R package to simulate a small regulatory network, and
- Introduction to High performance computing (HPC)
- HPC architectures
- Batch systems and using a Slurm scheduler
- HPC architectures
- How to scale up simulations on a HPC through profiling and optimisation
R for Genomics
Online
27 September
Register:
https://www.eventbrite.co.nz/e/introduction-to-r-for-genomics-online-tickets-415675064307
The focus of this workshop is on getting started with R while working with genomics data. Some of the topics covered in the workshop are:
- An introduction to R and RStudio
- R basics: The R language, reading data into R, storing data as objects
- R packages
- Tidyverse: easily manipulate and manage large datasets
- Publication-quality data presentation using ggplot2
- Knitr: keep track of workflow and produce easy-to-follow reports of your work
- Where to get more help when you are ready to do more
RNA-Seq Data Analysis
Online
02 November
Registration & session description coming soon...
Introduction to Bash Scripting and HPC Job Scheduler
Online
03 November
Registration & session description coming soon...
Otago Bioinformatics Spring School
Online
21 - 25 November
Registration has closed
Hosted at the University of Otago, the Bioinformatics Spring School 2022 is a week-long training event for researchers, supported by Genomics Aotearoa and the New Zealand eScience Infrastructure (NeSI). It will combine talks from researchers and hands on computational workshops (influenced by The Carpentries), covering example workflows of DNA variant calling, Genome assembly, environmental DNA (eDNA), and Nanopore. For more information, visit the Spring School website.
Metagenomics Summer School: Prokaryotic Metagenomics Workshop
In-person - University of Auckland
29 November - 02 December
Registration closes on 11 September:
https://auckland.au1.qualtrics.com/jfe/form/SV_eY6GW6gTWCNalo2
This is a comprehensive hands-on workshop featuring bioinformatics analysis, discussions, group assignments and talks. Topics include:
- Bash scripting
- Read quality filtering
- Prokaryotic metagenome assembly
- Assembly assessment and binning
- Prokaryotic and viral gene prediction and annotation
- Data analyses and visualisation





