Supporting ecological research through enhanced interpolation capability
The below case study shares some of the technical details and outcomes of the scientific and HPC-focused programming support provided to a research project through NeSI’s Consultancy Service.
This service supports projects across a range of domains, with an aim to lift researchers’ productivity, efficiency, and skills in research computing. If you are interested to learn more or apply for Consultancy support, visit our Consultancy Service page.
Research background
Ecological research such as species distribution modelling requires national scale maps of environmental variables. Often these data must be interpolated from point samples of environmental variables. Natural neighbour (or Sibson) interpolation provides a potentially useful technique as it is an exact interpolator – it creates a smooth surface free of any discontinuities. It is also a local method and is spatially adaptive. Moreover, it is possible to produce measures of uncertainty associated with the interpolation.
Project challenges
Tom Etherington, a Spatial Modeller with Manaaki Whenua - Landcare Research, has developed a Python code that applies Sibson/natural neighbour interpolation to ecological data distributed over New Zealand. While the code is easy to use, it was not scaling to the resolution required to cover New Zealand at the onset of the project. Tom asked NeSI Research Software Engineers to help him improve the model's performance. It was important for Tom to keep the code in Python for maintainability.
What was done
NeSI showed how to profile code to identify performance bottlenecks. With this step, NeSI found that small code changes can have a significant effect. For instance, moving query calls to the KD tree can be vectorised, reducing the execution time of this part of the code from 3 hours to 3 minutes.
NeSI implemented parallelism which further reduced the execution time. The sweet point was found to be about 10 processors providing a 5X speedup over the serial code.
Main outcomes
- Faster code, higher research productivity, a better designed software and a more efficient use of NeSI resources.
- Following the end of the project, the researcher discovered “numba”, a Python module that translates Python code to C without requiring the programmer to know C. This brought another performance improvement and shows that the efficiency of a code often continues to improve beyond the end of the project thanks to upskilling.
Researcher feedback
“I will now be able to make better maps of environmental variables, and hence much better models of species distributions. When it comes to writing efficient code I know very well I can only get so far on my own. Being able to get help from experts at NeSI such as Alex and Chris really moves my computational science onto another level. I can do my own limited ‘profiling’ by just running blocks of code separately, but getting a holistic view of where the bottlenecks are is incredibly useful – and something I must learn to do myself!"
- Tom Etherington, Spatial Modeller, Ecology, Manaaki Whenua - Landcare Research
The picture shows how natural neighbour interpolation works for a set of manufactured values.
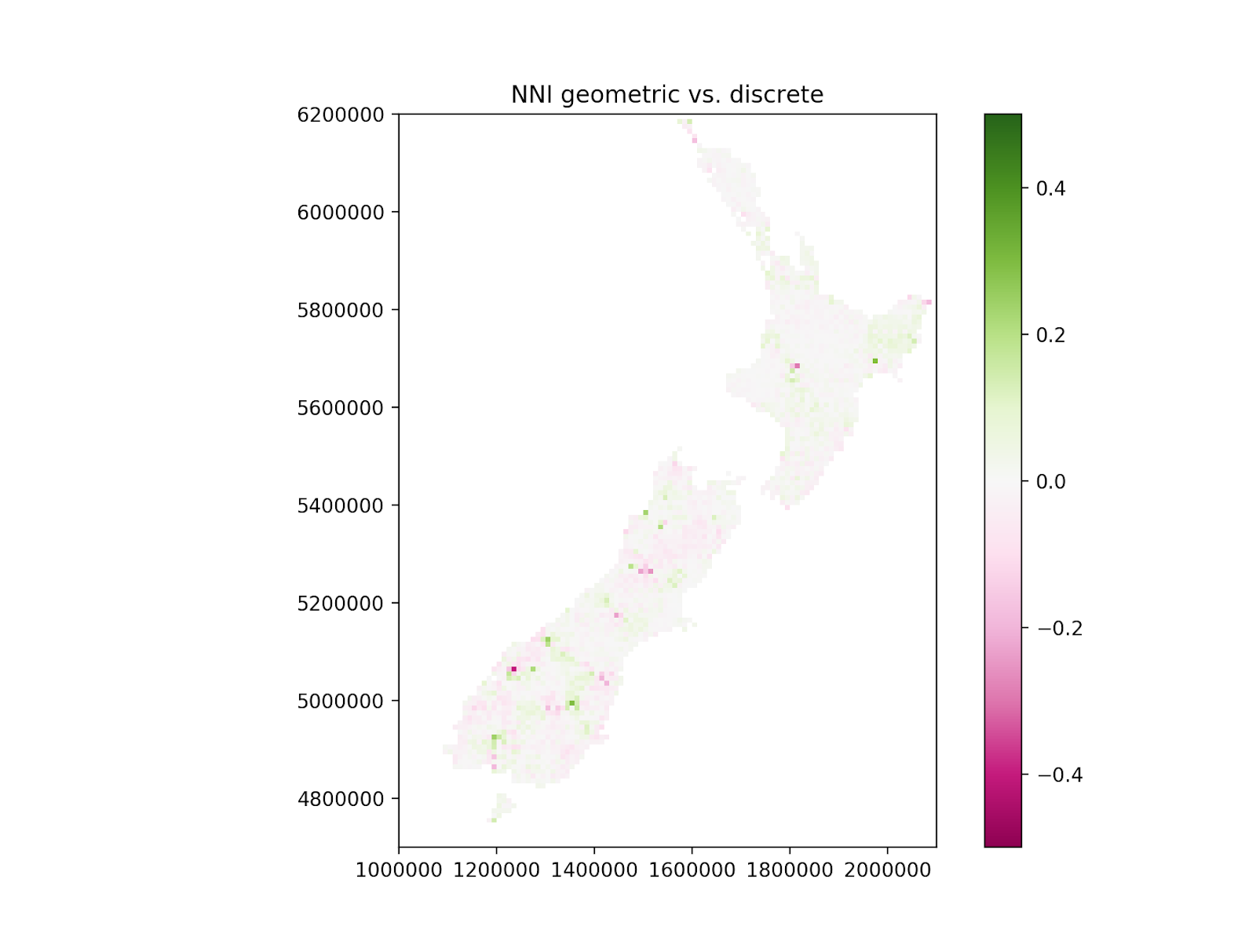
The circles are random sites for which field values were provided. The background colour represents the interpolated field values at pixel locations, which was computed from the field values at the sites
Do you have an research project that could benefit from working with NeSI research software engineers? Learn more about what kind of support they can offer and get in touch by emailing support@nesi.org.nz.






