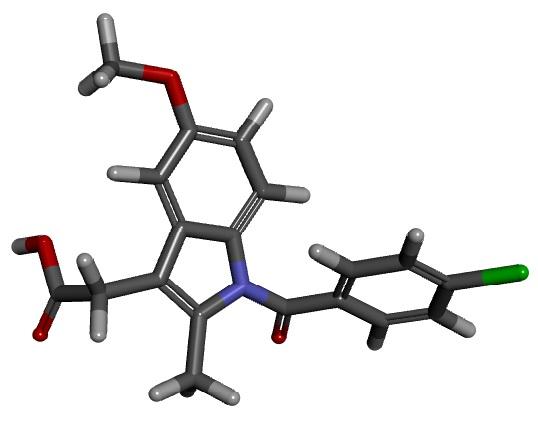
Predicting druglikeness in new compounds
Scientists in New Zealand and overseas are always on the hunt for new drugs, both naturally-occurring and synthetic, to treat illnesses and chronic conditions. In recent decades, as computers have become more powerful and more readily available, scientists have been relying more on computational methods such as computer-aided drug design to identify drug candidates. These techniques are quick and relatively inexpensive per compound, so when it comes time to synthesise drug candidates we can focus our efforts and resources on those compounds most likely to be viable.
Besides the drug candidate’s effectiveness in treating the target disease, there are other properties that interest us as we screen our libraries of compounds. These properties are often referred to collectively as “ADME-Tox”, which stands for Absorption, Distribution, Metabolism, Excretion and Toxicity. Even if a compound is shown to be extremely potent and effective in treating the disease, an unfavourable ADME-Tox profile means it is not a viable drug candidate. Perhaps the drug will just get flushed out of the body without being absorbed into the blood stream, or maybe it is absorbed too well and builds up in the body, or has unacceptably bad side effects.
When investigating a compound’s ADME-Tox profile, one of its key physical properties is the compound’s pKa, which is a measure of acidity. The lower a substance’s pKa, the greater that substance’s acidity (though a substance with a high pKa isn’t necessarily a strong base, just a very weak acid). Strongly acidic compounds tend to be highly water soluble but unable to pass easily through membranes such as the intestine. They also pose toxicity problems, as they disrupt the pH of the solution (e.g. partially digested food or blood plasma) and can damage delicate chemical structures in the body. On the other hand, very weak acids are often highly insoluble in water and so will have difficulty being absorbed into the body, transported to the correct places, or excreted once their work is done. The goal, therefore, is to identify those compounds that fit the desired pKa range for any given application.
While a compound’s pKa can be easily measured, this requires a physical sample which is often not available in the case of drug candidates that are being virtually screened. We are instead looking for a way to predict pKa values using computed data. It is not easy to accurately calculate pKa as some of the quantities involved, such as the energy needed to dissolve the compound in water, are difficult to compute with precision. However, one of the most significant components of a compound’s pKa is its proton affinity, which is the energy needed to remove the most weakly attached hydrogen atom from a single molecule of the compound. The proton affinities of most druglike molecules can be reliably computed using quantum-mechanical techniques. In this project, we aim to develop a model by which the pKa of a compound can be reliably predicted from its calculated proton affinity.
We were able to run some initial calculations of the proton affinities of small molecules on an ordinary personal computer, but a proper computation of a proton affinity requires quantum-mechanical approaches, demanding much greater computational resources as the size and complexity of the chemical system increases. The computational demands quickly grew to a point where we could only compute one proton affinity every few days, which made reliance on a desktop computer impractical. Once we started running our calculations on the NeSI clusters, we were able to greatly increase our throughput of new molecules as we could calculate the proton affinities of several molecules per day – a roughly 20-fold increase in research productivity.
We have also benefited from the expertise of NeSI’s Computational Science team. Before starting work on the cluster, Daniel had little experience with the Linux operating system and had never used a batch queueing system. We are now able to run calculations on many more molecules quickly and easily thanks to instruction and example workflows provided by NeSI staff, including tips for how to get the most out of computational chemistry software such as GAUSSIAN in a cluster environment.