Optimising tools used for genetic analysis
Knowledge of genetic relatedness is important in a number of genetic applications. Genetic relatedness estimates can be used to infer the relationship between individuals (e.g., mother-son, siblings) and can also be used by animal and plant breeders to predict the characteristics of offspring. Related to this, genetic linkage maps -- one-dimensional representations of genetic inheritance across the genome -- are also valuable as they form the starting point for a number of downstream analyses.
NeSI’s computational science team has been working with Timothy Bilton, a PhD student in the Department of Mathematics and Statistics at the University of Otago and at AgResearch, and Ken Dodds, a Statistician / Geneticist at AgResearch, to incorporate new computational methods and tools into their analysis of genetic relatedness and construction of genetic linkage maps.
“Both these applications can have high computing resource requirements with 10,000 to 1 million genomic positions and 100 – 100,000 individuals in a study, depending on the particular case,” says Ken. “Chris Scott and Wolfgang Hayek from NeSI helped our team profile our code to detect inefficiencies and helped improve our code, allowing us to run our analyses faster.”
In one test case, their code ran nearly 20 times faster (24 mins vs 8 hrs) after they implemented the code optimisation and computational tools recommended by NeSI.
Many of Timothy and Ken’s projects use genotyping-by-sequencing (GBS), a relatively new sequencing technology that enables many samples to be sequenced at high-throughput. The advantages of GBS over previous technology is that it is significantly cheaper and faster, and is easily adapted to a diverse range of species.
“GBS allows researchers to obtain genotype information across the genome of species that previously were not possible due to limited budgets, such as New Zealand native species,” says Timothy. “However, data generated by GBS contains additional errors that can severely affect the reliability of the results if they are not taken into account.”
Consequently, they need to use specialised software packages that can handle the nuances of GBS data in particular contexts. One of those is KGD, a package developed for estimating the genetic relatedness between individuals in a population. KGD can have an important role of performing quality control of GBS data, and a common application of KGD software is to check the accuracy of a recorded pedigree.
Another package, GUSMap, uses GBS data to construct genetic linkage maps. Errors in these maps can negatively impact the reliability of downstream analyses, so GUSMap is designed to handle any errors in the data and enable researchers to construct accurate linkage maps from GBS data at a significantly reduced cost compared to other sequencing technologies.
To improve Timothy and Ken’s productivity with these packages, NeSI introduced them to tools such as Rcpp, used for converting sections of R code into C++ for higher performance, and to OpenMP, an application programming interface for parallelisation (using multiple processors simultaneously to calculate results faster).
The support from NeSI has already benefited Timothy across multiple projects.
“Just recently, I managed to implement some OpenMP parallelization in another R package I’m working on, so that has been a big help,” he says. ““It would have taken a lot time to work out how to use Rcpp and OpenMP due to our limited programme knowledge. Chris and Wolfgang have saved us a great deal of time and pain trying to learn how to use these techniques to improve the efficiency of our code“
Recent Publication:
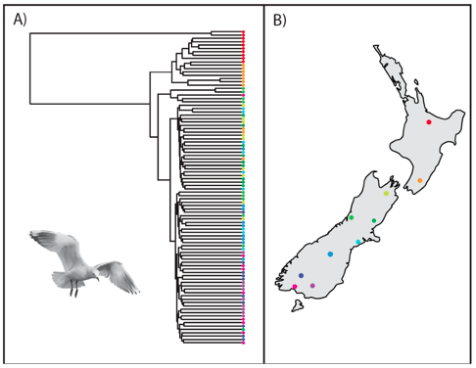
Mischler, C.; Veale, A.; van Stijn, T.; Brauning, R.; McEwan, J.C.; Maloney, R.; Robertson, B.C. Population Connectivity and Traces of Mitochondrial Introgression in New Zealand Black-Billed Gulls (Larus bulleri). Genes 2018, 9, 544. doi:10.3390/genes9110544
-----------------------------
Need some help with your research project? NeSI’s computational science team can assist with code optimisation, parallelisation, porting to GPUs, custom code development, and many other tasks. Email support@nesi.org.nz if you are interested to find out more or if you have a task you’d like NeSI to help you tackle.
-----------------------------
The Technical Details:
Below you’ll find a more technical description of the computational science support NeSI offered Timothy Bilton and Ken Dodds. If you have any questions about these details, contact support@nesi.org.nz.
Below is a summary of the main tools introduced to support the use of KGD and GUSMap code, and the key outcomes achieved.
KGD Code
- Showed how to use some profiling tools - Rprof and profvis
- Profiling the code showed it was already well optimised - no single function was taking a large proportion of time
- Important to build R against a parallel math library, such as Intel MKL, to take advantage of parallel tcrossprod and SVD functions
- Identified a number of functions that, when added together, took a significant proportion of run time
- Added parDist function, a parallel version of the dist function, in the heatmap call
- Converted other “kernels” of R code to C++ using the Rcpp library
- Overall improvement on the test dataset was ~1.8x speedup with these changes
- There is now a framework for converting more code to C++/OpenMP
GUSMap code
- Profiling showed most of the time was spent in C functions for computing likelihood and score (these functions are called by the two optimisation routines that are available in the code, EM and optim)
- Part of the code was already parallelised using parallel loops in R
- Smaller test case
- Converted likelihood and score C functions to use OpenMP parallelisation
- Provided a detailed explanation of how we implemented OpenMP parallelisation
- EM version ~16x faster (36 cores)
- Optim version ~10x faster (36 cores)
- Larger test case
- Identified some matrix calculations that could be moved to C and parallelised with OpenMP (in addition to the likelihood calculation also being parallelised)
- Overall 20x faster on 16 cores (24 mins vs 8 hrs)







