Machine learning for clinical decision making
The below case study shares some of the technical details and outcomes of the scientific and HPC-focused programming support provided to a research project through NeSI’s Consultancy Service.
This service supports projects across a range of domains, with an aim to lift researchers’ productivity, efficiency, and skills in research computing. If you are interested to learn more or apply for Consultancy support, visit our Consultancy Service page.
Research background
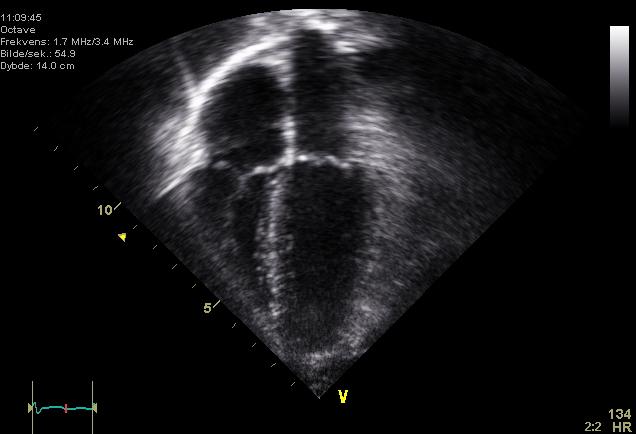
Nathan Russell, a University of Otago PhD student, is studying how to improve the use of "big data" in clinical decision making. He's exploring how machine learning tools can be leveraged to improve data integration, analysis, and visualisation. In this phase of his project, Nathan wanted to evaluate deep learning models for echo-cardiography images segmentation. His aim is to identify if available models are sufficient or if dedicated models should be developed in the future.
Project challenges
Nathan wanted to use a publicly available toolbox, EchoCV (rahuldeo/echocv ), a computer vision toolbox that supports echo-cardiography images view classification and segmentation using deep learning. However, the code relies on deprecated technologies (Python 2.7 and Tensorflow v1) and has not been maintained during the last two years to adapt it to newer versions of its dependencies. Therefore it needed adaptations to run on Mahuika.
What was done
As part of this Consultancy project, NeSI research software engineers installed the toolbox on Mahuika, ensuring the compatibility of the dependencies with the HPC GPUs. Changes in the code were made to ensure that the new data could be used, avoiding hard coded paths that would prevent re-usability of the tool. The refactoring also removed complexity from the data loading to the inference stage, to reduce the risk of bugs due to unnecessary redundant steps (e.g. nested conversions of input images using several tools when it can be achieved in one function call). Additional options were added to ensure Nathan can adapt his dataset to as close as possible to the original dataset used to train the neural network. These preprocessing options helped to diagnose the impact of minor changes in the input data on the final segmentation.
Main outcomes
The code is now capable of running on Mahuika.
The code can now use NeSI’s P100 GPUs, which offer faster processing power than CPUs, to run models from EchoCV.
With the EchoCV code adapted to run on Mahuika, Nathan was able to process his dataset on NeSI's HPC platform.
Nathan became more familiar with using Jupyter on NeSI, and was able to upskill in version control and command line usage, and developing Python coding skills for image segmentation.
Researcher Feedback
“Through the Consultancy with NeSI (Maxime), I was able to develop my code to be far more efficient, not only in its image segmentation task but also in its utilization of hardware, going from seven-minute runs on the CPU to under one minute on Mahuika's GPU cluster."
- Nathan Russell, PhD student, Bioinformatics and clinical data analysis, University of Otago and ESR
Do you have a research project that could benefit from working with NeSI research software engineers or a data engineer? Learn more about what kind of support they can offer and get in touch by emailing support@nesi.org.nz.